Can You Image That? Imaging a Cell and Its Proteins Together
Observing the world around us is a natural human instinct, and exploring the realm of the tiny and beautiful is especially captivating for scientists and the public alike. The business of building and testing new microscopes, and developing new methods of microscopy, is rapidly changing and evolving over time. As early as 1914, scientists started documenting the history of the microscope.
Thankfully, rather than drawing out what we see through the lens onto a piece of paper, there are now advanced forms of microscopy, like electron microscopy, that allow scientists to scan a substance—with an electron beam—to sample its topography, or surface shape, and produce wonderfully detailed images that also contain tremendous amounts of data. In a separate form of microscopy called fluorescence microscopy, fluorescent proteins—proteins in the cell engineered to have a fluorescent tag—and intracellular proteins fused to a fluorescent protein can be imaged because they emit light in response to certain wavelengths of light coming from the microscope.
In a continuous effort to “see what’s going on a bit better,” Howard Hughes Medical Institute researchers published an article in PLOS ONE today detailing improvements of their relatively new form of 3D super-resolution microscopy. It combines what they call PhotoActivated Localization Microscopy, or PALM (a type of fluorescence microscopy), with electron microscopy (EM), and lets us have a look at organelles, like mitochondria, and the location of nearby proteins, right in the same cell at roughly the same time.
For instance, if we know a specific protein attaches directly to DNA in a specific organelle, PALM allows us to see precisely the nanometer-scale location of this protein, when it is fused to a fluorescent protein. EM on the other hand, allows us to see the overall structure of the organelle, and so combining these to see both at the same time is extremely useful to cell biologists studying structure and function.
In their study design, PALM needs to be performed before EM (see image of sample preparation and setup below), and then the two images are overlaid by correlating the area of fluorescence, seen during fluorescence microscopy, to the area of the cell structure seen during electron microscopy. The overlaid image is the final result, and the goals of the researchers in this improvement study were to optimize the image resolution and the number of useable fluorescent dyes, speed up the protocol, and simplify the equipment involved by moving it from 3D (very difficult, less accessible) to 2D (easier, common in research institutions), in hopes of making this technique accessible to cell biologists.
In this and the previous work, the researchers describe how they combined the two forms of microscopy to achieve the results they were looking for. All forms of microscopy have limitations, especially when it comes to the limits of the optics and sample preparation, and in this case, scientists overcame a barrier between optimizing the available fluorescence and also optimizing the quality of the EM images that were produced.
As you might imagine, the better the overall image quality is, the better biologists can use the image information to help understand the structure and function of biological components, such as organelles and proteins. Additionally, though the cells used in this study were frozen and prepared, there is a possibility that live cells could be directly imaged using this technique.
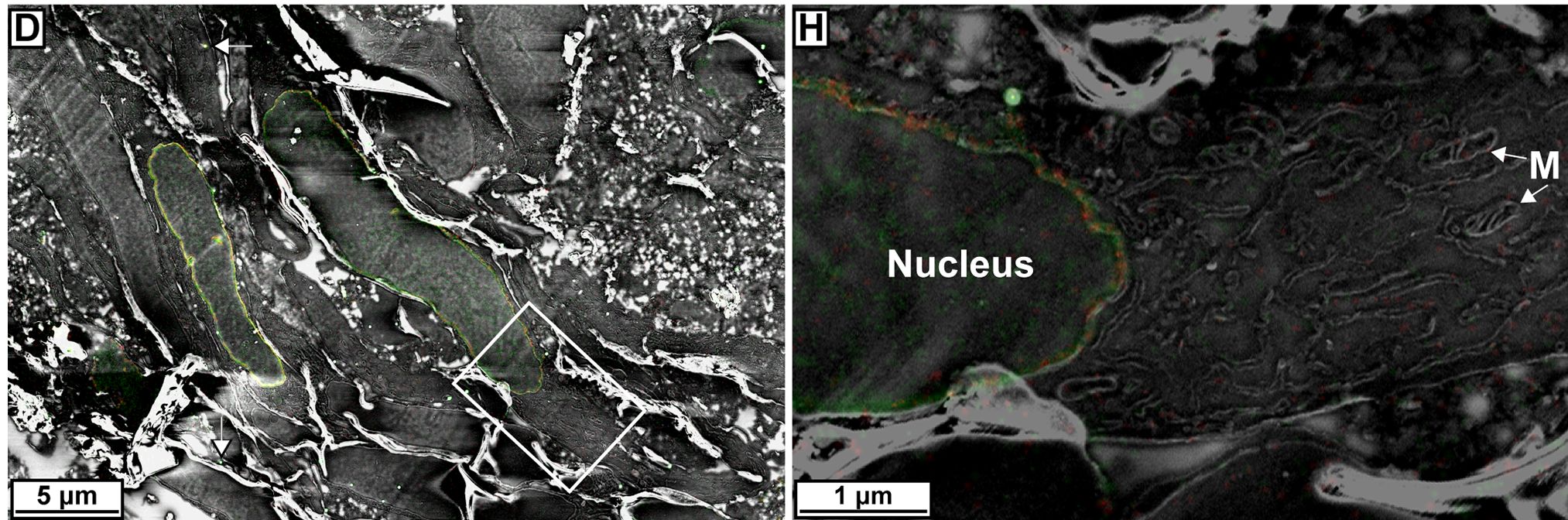
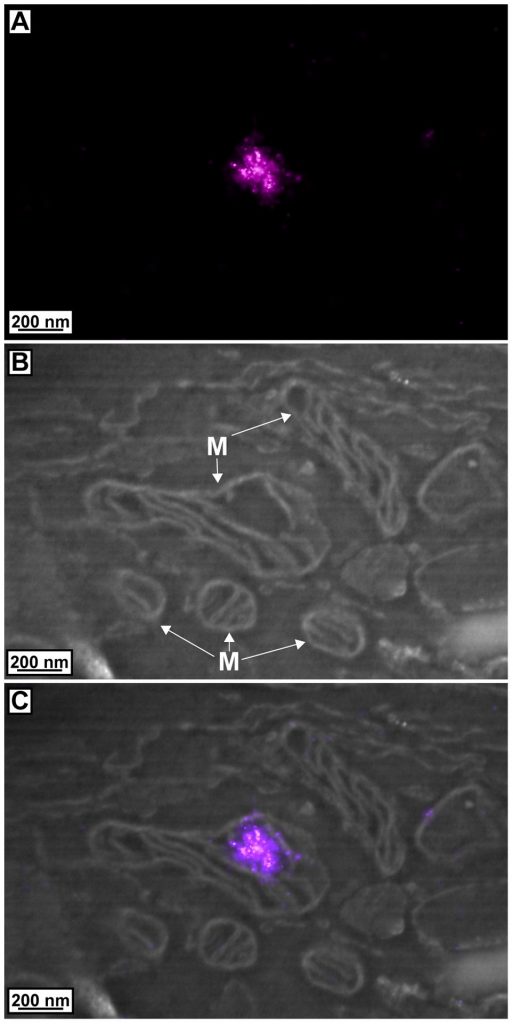
Though explaining and understanding the method are a little complicated, the pictures make it all worth it—and the scientists would agree. Below are example image sets. The first set is of an image containing mitochondria, or the powerhouse of the cell, and a protein that localizes in its DNA.
The first image in the set shows the result of PALM, the second contains the EM, and the third is the alignment of the two images (and M is for mitochondria).
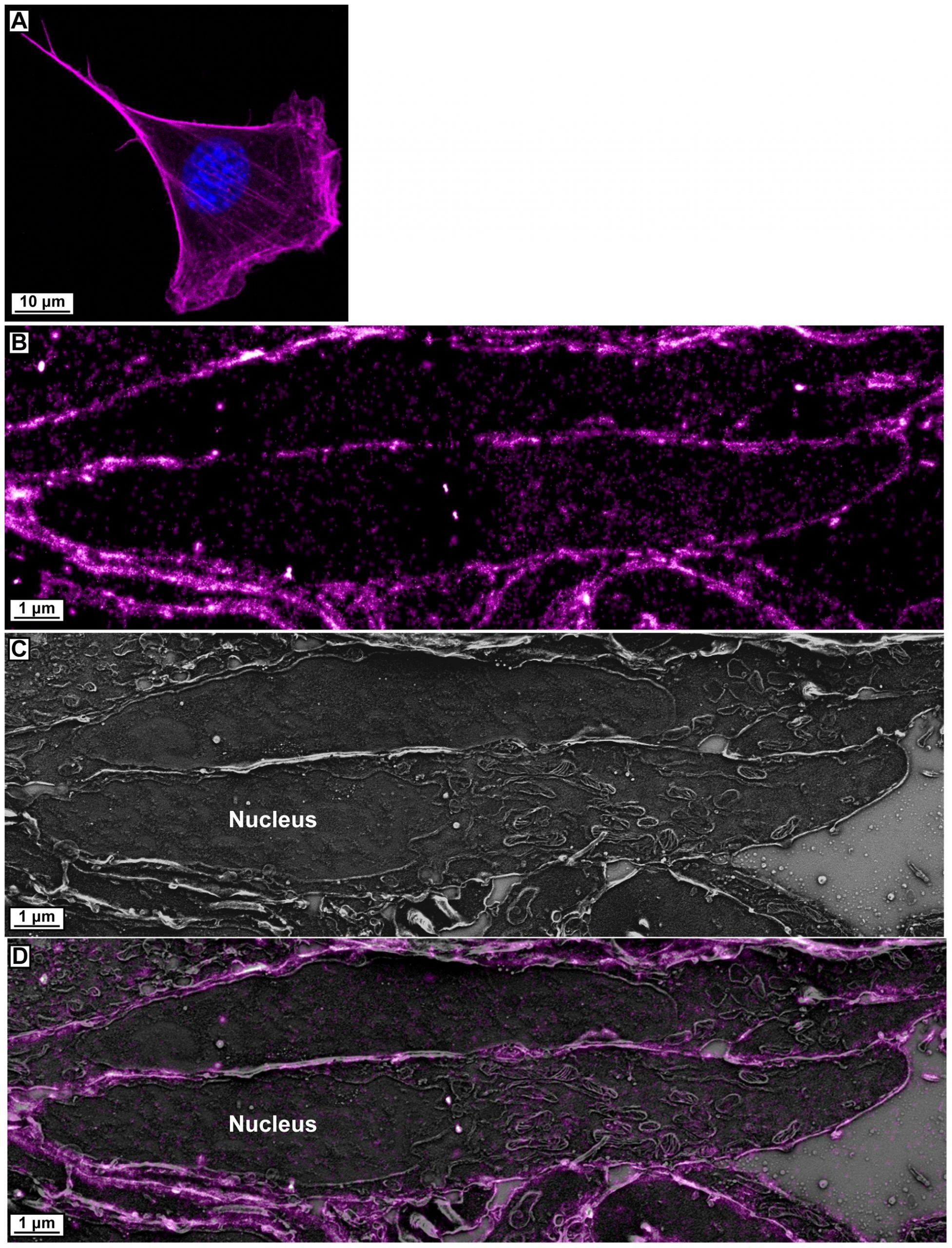
And here’s a second set, with imaging of a whole cell by traditional confocal microscopy (A), followed by a similar sequence as above (B-D) for a nucleus and the location of a fluorescent dye that adheres to actin (protein) filaments.
The researchers also managed to perform imaging using two colors (title image), demonstrating the versatility of the technique. Check out the full details and image set here, as well as other recent studies involving new imaging techniques here, here, and here.
Citations:
Gaietta GM, Giepmans BNG, Deerinck TJ, Smith WB, Ngan L, Llopis J, Adams SR, Tsien RY, and Ellisman MH (2006) Golgi twins in late mitosis revealed by genetically encoded tags for live cell imaging and correlated electron microscopy. PNAS 103(47) doi:10.1073/pnas.0608509103
Kopek BG, Shtengel G, Grimm JB, Clayton DA, Hess HF (2013) Correlative Photoactivated Localization and Scanning Electron Microscopy. PLoS ONE 8(10): e77209. doi:10.1371/journal.pone.0077209
Kopek BG, Shtengel G, Xu CS, Hess HF (2012) Correlative Photoactivated Localization and Scanning Electron Microscopy. PNAS Early Edition. doi:10.1073/pnas.1121558109
Image Credits: All images from the PLOS ONE article doi:10.1371/journal.pone.0077209